Preparative
Published over 6 years ago. See the latest and most current information on Preparative.
Scientist’s understanding of the biological world has increased exponentially over the last few years, resulting in the development of a whole new range of pharmaceuticals based on proteins rather than the traditional small molecules that the pharmaceutical industry was associated with [1]. With the increase in attention associated with the development of protein-based therapeutics, there is also a need for analytical science to support this development by ensuring that appropriate technology is being used to characterise the new products to ensure that the drug is efficacious and nontoxic. This has resulted in the development of new workflows allowing the characterisation of protein therapeutics but has also created new analytical challenges that are not associated with the analysis of small molecules [2].
Within the field of chromatography, one of the fundamental drivers for change has been the need to decrease the analysis time. One approach that has been very successfully employed that ensures that this can be achieved is the application of coreshell particles and smaller (<2 μm) particles and the development of UHPLC. For small molecules this has been shown to be very effective and has had a transformational effect in many QC laboratories reducing analysis times, solvent consumption and most importantly improving the sensitivity of the assays. The same technology has also started to be applied to the analysis of larger, protein-based molecule, however challenges have arisen due to the complexity of the molecules that are being analysed. One such area where the use of sub 2 μm has been applied to is in size exclusion chromatography.
Size exclusion chromatography is an important part of the workflow for characterising proteins, and specifically mAbs, as it is the only chromatographic technique which can be used for the determination of aggregates that keeps the protein in its native state. Other chromatographic techniques either rely on;
• a chemistry driven separation mechanism and consequently it is potentially harder to differentiate molecules that may have the same level of chemical interaction but have a different mass due to the formation of aggregates.
• Operating in an environment which disrupts the structure of the protein through the use of a chaotropic solvent, such as acetonitrile.
Size exclusion chromatography separates based on the hydrodynamic radius of a molecule. The concept was first postulated by Synge and Tiselius [3] when investigating the properties of zeolites, with the first examples being demonstrated by Wheaton and Bauman [4], and the first application to the analysis of proteins being demonstrated by Lindqvist and Storgårds [5]. There are many names that are given to this process sized based separation process; gel permeation chromatography (GPC) is common when using organic solvent as the mobile phase with a hydrophobic stationary phase, while gel filtration chromatography (GFC) [6] or more recently size exclusion chromatography (SEC) [7] is used for separations in an aqueous mobile phase with a hydrophilic stationary phase.
In this case it has been shown that the retention time for an individual analyte is directly proportional to the log of the relative molecular mass (M) [8], for molecules that are neither completely excluded or for molecules that can penetrate all of the pore network, (eqn. 1).
Eqn. 1
Where;
m and b are the slope and intercept of the linear part of the calibration line, and KD, the thermodynamic retention factor, is given by the following expression (eqn. 2);
Eqn 2.
Where;
VR – retention volume of the analyte/protein
V0 – retention volume of the column
Vtot –total solvent volume of the column
While theory predicts a perfectly linear correlation, the variability in exact pore structure often leads to a degree of nonlinearity. Calibrations are run using a series of known molecular weight samples to allow identification of precise retention times for that particular molecular weight (MW). Thereby when determining the molecular mass of unknowns, comparison of retention times with known standards will give a good indication of the size of the unknown molecule. The calibration curve provides an upper and lower MW limit that the column is able to separate, where a sharp upward and downward knee forms in the calibration curve.
The formation of aggregates can have substantial impact on the toxicity of the therapeutic protein [9] and it is therefore important to be able to measure the amount of aggregation either for final product quality, but also as an in-process measurement to ensure product quality. Thus, there is a need for fast aggregate analysis, which would seem to lend itself to UHPLC and also potentially running the assay at elevated temperature to allow for faster flow rates to be used.
The introduction of UHPLC [10] into the separation scientist’s vocabulary has allowed for the substantial reduction in analysis times, some of which can be attributed to the offsetting of the higher resolution capabilities of the smaller stationary phase particles and some of which can be attributed to the redevelopment of the assay using more controlled surfaces available on the newer column formats. It is therefore an obvious development to apply the learnings associated with small molecule analysis into the field of size exclusion chromatography for the analysis of large protein-based molecules. Small molecule therapeutics are chemically synthesised, as opposed to the genetically engineered proteins derived from living cells that are used in biotherapeutics.
Small molecular therapeutics are chemically synthesised, compared to the genetically engineered proteins used as biotherapeutics. Biopharmaceuticals may be produced from microbial cells (e.g., recombinant E. coli or yeast cultures), mammalian cell lines, plant cell cultures, and moss plants in bioreactors of various configurations, including photo-bioreactors. This potentially allows for the opportunity to have virtually direct measurement of the quality of the protein broth to ensure product quality is maintained.
This concept has resulted in manufacturers developing a new generation of UHPLC SEC stationary phases. This is not a trivial task in itself, as the pore size required to generate a separation of molecules having a molecular mass of ca. 150 kDa, substantially decreases the strength of the particle, with a consequent effect on lifetime and also column performance as the particle breaks due to the increase in pressure and also the lack of particle strength to cope with the increase in pressure. There is however a degree of caution that needs to be taken when using higher pressure chromatographic systems with proteins. The ultimate aim is to analyse the sample in its natural form, with little or no changes affecting the structure of the protein. An increase in pressure can result in deformation of the protein, this can affect the degree of aggregation in the sample, as aggregation can be deemed to be an equilibrium process which will be affected by the external pressure. Thus, increasing the pressure can result in an increase in the aggregated form of the molecule [11].
As well as potentially affecting the shape directly, increasing the pressure drop across the column can have an impact on the temperature that the proteins will experience. So consequently, care has to be taken that excessive pressures are not being employed which would result in the possibility of thermally induced protein denaturation [12].
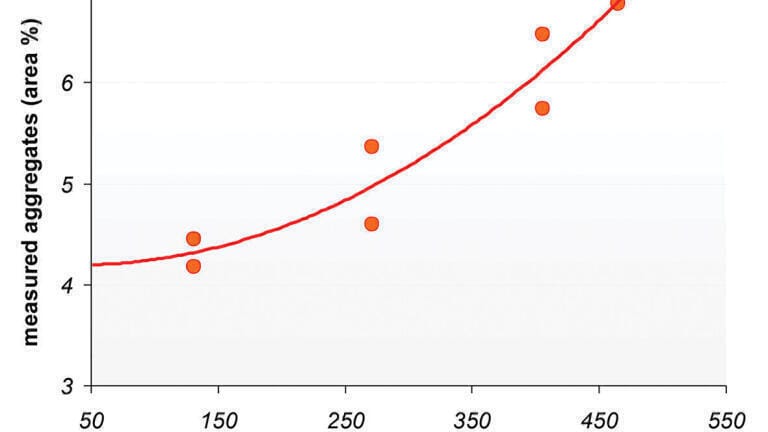
In order to test out this hypothesis, researchers at the University of Geneva [8] performed a series of experiments where the pressure applied to the SEC column was altered by the use of a series of linear flow restrictors which resulted in the following column head pressures, 131, 271, 406 and 465 bar.
Column: Acquity UPLC BEH200 SEC 1.7 μm, 150 mm × 4.6 mm.
Mobile phase: 20 mM disodium hydrogen-phosphate buffer of pH = 6.8.
Flow rate: 200 µl/min
Detection: FL (Ex: 280 nm, Em: 360 nm).
Sample: Heat stressed panitumumab
Figure 2 shows two examples from the research performed by Fekete et al. [13] that demonstrate the effect that pressure has on the protein aggregation. It can be clearly seen that increasing the pressure results in an increase in the aggregated form of the panitumumab. Increasing the pressure from 131 bar to over 400 bar results in the measured aggregation by more than 50%. It can also be observed that there is a very distinctive correlation between the amount of observed aggregation and the column head pressure. Clearly this would be very impactful on a separation scientist attempting to determine a critical quality attribute associated with the therapeutic protein product [14-15].
UHPLC has had a significant benefit for separation scientists. The impact in terms of improved resolution has resulted in laboratories being much more efficient in sample turn-around. It has resulted in a better understanding of the chemical processes that are being monitored, whether this is a man-made synthetic process or if it is a better definition of a complex biologically derived sample. However, as with all measurement science it is important to be aware of the impact that the measurement system can have on the determination of the compounds under investigation. The use of SEC at elevated pressures has been demonstrated to impact on the nature of the sample and although it could be reasonably anticipated that this effect is very dependent on the protein under investigation, the separation scientist should be aware of the impact that the pressure can have on the determination of aggregates. It is suggested that performing a SEC separation of proteins at pressures below 100 bar should ensure that there are no detrimental effects on the nature of the shape of the protein, which could result in an inaccurate determination of a critical quality attribute. Proteins, just like the scientists analysing them, under pressure may not behave in a natural manner.
1. G. Walsh, Biopharmaceutical benchmarks 2018, Nature Biotechnology volume36, pages1136–1145 (2018)
2. A. B. Chakraborty, H. Xie, St John Skilton, J. C. Gebler, W. Chen, LCGC Volume 7, Issue 3, pg 26–33
3. R. Synge, A. Tiselius, Biochemical J., 46 (1950) xli
4. R.M. Wheaton, W.C. Bauman, Annals of the New York Academy of Sciences 57(3) (1953) 159-176
5. B. Lindqvist, T. Storgårds, Nature 175 (1955) 511-512
6. K C. Duong-Ly, S B. Gabelli, Methods in Enzymology, 541 (2014) 105-114
7. K. Stulk, V. Pacakova, M. Ticha, J. Biochem. Biophys. Met. 56 (2003) 1–13
8. M. Dole, L. Mach, R. L. Hines, R. C. Mobley, L. D. Ferguson, M. B. Alice, L.L. Mack, L. L. Mack , J. Chem. Phys., 49 (1968) 2240-2249
9. E. M. Moussa, J. P. Panchal, B. S. Moorthy, J. S. Blum, M. K. Joubert, L. O. Narhi, E. M. Topp, Journal of Pharmaceutical Sciences 105 (2016) 417e430
10. R. Houghton, P. Grace, Chromatography Today, Feb. 2008, 5-7.
11. R.F. Tilton Jr, J.C. Dewan, G.A. Petsko, Biochemistry 1992, 31, 9, 2469-248
12. V. V. Mozhaev, K. Heremans, J. Frank, P. Masson, C. Balny, PROTEINS: Structure, Function, and Genetics 24:81-91 (1996)
13. S. Fekete, K. Ganzler, D. Guillarme, J. Pharm. Biomed. Anal. 78–79 (2013) 141-149
14. S. Fekete, J.L. Veuthey, D.V. McCalleym J. Chromatogr A. 2012 Dec 28;1270:127-38
15. J. De Vos, E.R. Kaal, R. Swart, M. Baca, J. Sep. Sci. 2016 Feb;39(4):689-95
