Electrophoretic separations
Published over 7 years ago. See the latest and most current information on Electrophoretic separations.
To address biomedical questions intrinsically dealing with limited sample amounts with a metabolomics approach, the design of novel or improved analytical techniques is needed. In this paper, we highlight the possibilities of capillary electrophoresis-mass spectrometry (CE-MS) as a microscale analytical platform for metabolic profiling of small amounts of biological material. A few examples from my group are discussed in order to show the utility of CE-MS for this purpose. Finally, we reflect on a number of analytical challenges that still need to be addressed in order to make CE-MS a viable approach for material-limited metabolomics.
Metabolomics has become an important tool in biomedical and clinical research and is particularly promising in biomarker discovery. Currently, reversed-phase LC-MS, GC-MS and nuclear magnetic resonance spectroscopy (NMR) are used as the main analytical tools for metabolomics studies. These analytical techniques require sample material which often ranges in the amount from 10 to 50 μL for LC-MS and GC-MS (in this case needed for both sample preparation and injection), and up to 500 μL for NMR. Consequently, the need for these minimal sample amounts often prevents their use for biomedical and clinical problems inherently dealing with very low amounts of material [1-3]. For example, metabolic profiling of cerebrospinal fluid (CSF) from transgenic mouse models, spheroids/microtissues, liquid biopsies and samples from microfluidic 3D cell culture models, is seriously hindered by the standard analytical technologies.
Until now, relatively little effort has been made to downscale the analytical technique and /or workflow for metabolomics studies, as the application has been often focused on addressing biomedical and clinical questions associated with human urine and plasma (or serum) samples. In order to enable micro-metabolomics, i.e. the metabolomics study of biological and biomedical questions intrinsically dealing with small amounts of sample material, new analytical technology with a dramatically improved sensitivity is needed. The approach of micro-metabolomics was first coined by Moco et al. in the field of plant metabolomics [4], however, here we refer to this approach in the context of biomedical and clinical questions. It is our conviction that the successful development of a microscale analytical platform for material-restricted biological samples will constitute a real breakthrough in the field of metabolomics as it will open the way for a deeper understanding of biological processes in sample-limited cases.
Capillary electrophoresis-mass spectrometry (CE-MS) can be considered an attractive microscale analytical technique for addressing biological questions inherently dealing with low amounts of material [5-10]. An example that cannot be properly studied with the current standard analytical tools, and which is of particular interest to our group, concerns the disentanglement of the behaviour of a single cell within a group or population of mammalian cells, and as such to obtain a better understanding on the role of cell heterogeneity in tumour biology.
In CE, nanolitre injection volumes can be used from just a few microliters of sample or less in the injection vial. Therefore, CE-MS can be regarded as a highly suited approach for the analysis of especially polar and charged metabolites in tiny sample amounts, as demonstrated by our group for example for mouse CSF [11]. This body fluid can only be obtained in a few microliters under the right experimental conditions. By employing a 1:1 dilution of CSF with water, and as a result completely retaining sample integrity, more than 300 compounds could be detected by CE-MS. For the injection only 45 nL of the sample was consumed from a vial containing not more than 2 µL of a 1:1 diluted CSF. Therefore, the CE-MS approach allows multiple analyses on a single highly valuable mouse CSF sample, enabling repeatability studies and, even more interestingly, to analyse the same sample at different separation conditions in order to enhance resolution and therefore metabolic coverage. The ability to perform multiple analyses on a single scarcely available biological sample is not possible with the conventional analytical techniques employed in metabolomics.
In CE, analytes are separated on the basis of differences in their intrinsic electrophoretic mobilities which is, except for the low sample and solvent requirement of CE, fundamentally different from chromatographic-based analytical techniques. Consequently, CE-MS provides a complementary view on the composition of endogenous metabolites present in a given biological sample [12]. Compared to chromatographic-based separation methods the separation efficiency of CE is very high due to the flat flow profile of the electro-osmotic flow. Moreover, there is no mass transfer between phases and therefore only longitudinal diffusion contributes to band broadening under proper experimental conditions. The intrinsically high separation efficiency of CE is very advantageous for the high resolution separation of structurally similar metabolites in complex samples.
Until now, various research groups have developed CE-MS approaches for metabolic profiling of limited sample amounts, and also for single cell analysis [3, 10]. Concerning the latter, the metabolomics studies were often focused on the analysis of a relatively large single non-mammalian cell (diameter in the range of 100 to 1000 µm with a cellular sample content ranging from circa 10 to 1000 nL). The content of a single mammalian cell is estimated to be around 1 pL (diameter ~10 µm) [13], and that of a single HepG2 cell is estimated to be around 3 pL (diameter ~12 µm). Therefore, the profiling of (endogenous) metabolites in a single mammalian cell is clearly a vast analytical challenge. Over the past few years, various research groups have done pioneering work in the development of analytical techniques for acquiring metabolic profiles from a single cell [14-19]. For an overview of these technological developments, we refer to the following recent reviews [20-22].
In my group, we have recently assessed the utility of CE-MS employing a sheathless porous tip interface for metabolic profiling of low numbers of mammalian cells using HepG2 cells as a model system [23]. The aim was to profile a wide range of endogenous metabolites in just a few cells, as the latter will enable to study the effect of cell heterogeneity, which really matters in various key fundamental biological questions. The sheathless porous tip interface was developed by Moini a decade ago by removing the polyimide coating of a fused-silica capillary outlet via etching of the capillary wall with 49% solution of hydrofluoric acid to a thickness of about 5 μm [24]. The electrical connection to the capillary outlet is achieved in this configuration by inserting the etched conductor into an ESI needle which is filled with separation buffer. The sheathless porous tip design is especially useful for interfacing narrow (<30 μm inner diameter) capillaries and for low flow-rate (<100 nL/min) nano-ESI-MS analyses.
By the integration of an in-capillary preconcentration technique, in this case transient isotachophoresis, in the sheathless CE-MS approach low nanomolar detection limits could be easily obtained for a wide range of basic metabolites (e.g., amino acids, amines, small peptides). It is important to stress that low nanomolar detection limits could be obtained by only using an injection volume of circa 40 nL (total capillary volume is ~600 nL). In cases where sample volume is not an issue LC-MS will be a preferred analytical technique, however, it is important to highlight that reversed-phase LC-MS is not a suitable tool for the analysis of highly polar and charged metabolites.
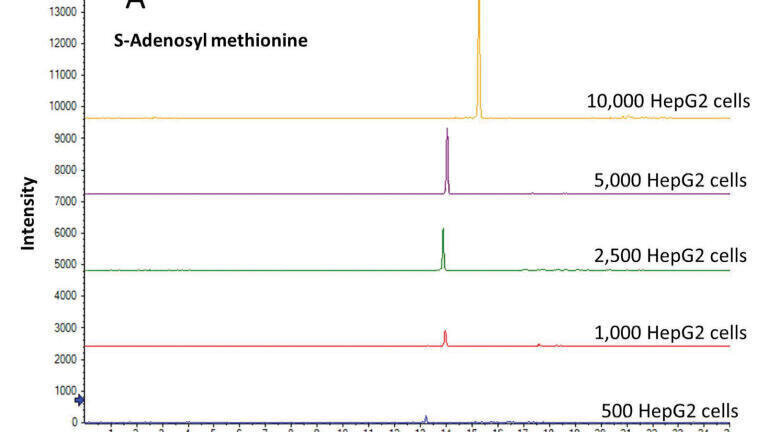
Given the nanomolar concentration sensitivity, the sheathless CE-MS method could be effectively used for acquiring metabolic profiles in HepG2 cells starting from 10,000 down to 500 cells, which corresponds to the injection content of 5 HepG2 cells to less than one cell. These results suggest that the method has the sensitivity for performing single cell mammalian metabolomics studies. Though, the sheathless CE-MS method has been primarily used as a screening method so far, we have also compared the detector response (using peak area as read-out) for a few selected endogenous metabolites as a function of the starting amount of HepG2 cells. As shown in Figure 1, this analysis revealed a linear detector response (R2=0.9986 and 0.9948 for S-adenosyl methionine and glutamic acid, respectively) when going from 10,000 to 500 HepG2 cells, indicating the potential of the sheathless CE-MS method for quantitative metabolomics studies of limited sample amounts. The performance of the approach in terms of repeatability for profiling metabolites in small amounts of HepG2 cells was assessed by the consecutive analyses of a single extract from 10,000 HepG2 cells. The relative standard deviation (RSD) values for migration times and peak areas of selected endogenous metabolites were below 3% and 10%, respectively, which are acceptable values for studies employed under ultra-low flow-rate separation conditions in conjunction with nano-ESI-MS conditions.
Metabolomics studies dealing with low amounts of biological sample have to critically consider pre-analytical steps as adsorption effects (for example, to sample vials, pipette tips, etc.), notably with sample volumes far below 1 µL, may result in significant analyte losses. A more critical factor is the fraction of the sample that needs to be (effectively) injected into the CE-MS system. In this context, miniaturisation and optimisation of sample preparation will be crucial for material-limited metabolomics studies. For CE-MS-based metabolomics studies, fractionation procedures based on liquid-liquid extraction for the selective isolation of polar and charged metabolites (which obviously will be in the polar phase), from proteins and non-polar compounds in small amounts of biological samples can be considered. The polar phase/fraction can be further processed using enrichment tools as electroextraction or depleted zone isotachophoresis [25-27]. For example, in electroextraction, analytes can be extracted from a donor phase, i.e. the sample, into an acceptor phase by using an electric field. Alternatively, polar and charged metabolites can be selectively enriched from a biological sample via an organic layer into a hanging aqueous droplet prior to further analysis. With depletion zone isotachophoresis, charged compounds can be preconcentrated and separated in a microfluidic channel according to their electrophoretic mobilities, adjacent to a depleted zone created by a nanochannel. Afterwards, the charged compounds can be released in fractions according to their electrophoretic mobility for further analysis. In the proposed sample preparation steps, the fractionated polar and charged compounds can be further separated by the sheathless CE-MS method.
Overall, the combination of an effective, tailor-made sample preparation method with sheathless CE-MS will enable us to acquire metabolic profiles from a low number of mammalian cells, ultimately a single mammalian cell, in a robust way. We anticipate that such a fully optimised microscale analytical workflow will be of high value to nanodosing studies and single cell biopsies, especially in the context of cancer research.
Wei Zhang would like to acknowledge the China Scholarship Council (CSC, No. 201507060011). Rawi Ramautar would like to acknowledge the financial support of the Vidi grant scheme of the Netherlands Organization for Scientific Research (NWO Vidi 723.016.003).
Conflict of Interest
The authors have no other relevant affiliations or financial involvement with any organisation or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
1. Fessenden, M., Nature 2016, 540, 153-155.
2. Miggiels, P., Wouters, B., van Westen, G. J. P., Dubbelman, A.-C., Hankemeier, T., TrAC Trends in Analytical Chemistry 2018.
3. DeLaney, K., Sauer, C. S., Vu, N. Q., Li, L., Molecules 2018, 24.
4. Moco, S., Schneider, B., Vervoort, J., J Proteome Res 2009, 8, 1694-1703.
5. Ramautar, R., Lc Gc Eur 2017, 30, 658-661.
6. Zhang, W., Hankemeier, T., Ramautar, R., Curr Opin Biotechnol 2017, 43, 1-7.
7. Nemes, P., Knolhoff, A. M., Rubakhin, S. S., Sweedler, J. V., ACS Chem Neurosci 2012, 3, 782-792.
8. Onjiko, R. M., Portero, E. P., Moody, S. A., Nemes, P., J Vis Exp 2017.
9. Lapainis, T., Rubakhin, S. S., Sweedler, J. V., Anal Chem 2009, 81, 5858-5864.
10. Onjiko, R. M., Portero, E. P., Nemes, P., Capillary Electrophoresis–Mass Spectrometry for Metabolomics, The Royal Society of Chemistry 2018, pp. 209-224.
11. Ramautar, R., Shyti, R., Schoenmaker, B., de Groote, L., Derks, R. J. E., Ferrari, M. D., van den Maagdenberg, A. M. J. M., Deelder, A. M., Mayboroda, O. A., Anal Bioanal Chem 2012, 404, 2895-2900.
12. Godzien, J., Garcia, A., López-Gonzalvez, A., Barbas, C., Capillary Electrophoresis–Mass Spectrometry for Metabolomics, The Royal Society of Chemistry 2018, pp. 161-183.
13. Zenobi, R., Science 2013, 342, 1243259.
14. Guillaume-Gentil, O., Rey, T., Kiefer, P., Ibanez, A. J., Steinhoff, R., Bronnimann, R., Dorwling-Carter, L., Zambelli, T., Zenobi, R., Vorholt, J. A., Anal Chem 2017, 89, 5017-5023.
15. Abouleila, Y., Onidani, K., Ali, A., Shoji, H., Kawai, T., Lim, C. T., Kumar, V., Okaya, S., Kato, K., Hiyama, E., Yanagida, T., Masujima, T., Shimizu, Y., Honda, K., Cancer Sci 2018.
16. Masuda, K., Abouleila, Y., Ali, A., Yanagida, T., Masujima, T., Methods Mol Biol 2018, 1778, 269-282.
17. Aerts, J. T., Louis, K. R., Crandall, S. R., Govindaiah, G., Cox, C. L., Sweedler, J. V., Anal Chem 2014, 86, 3203-3208.
18. Zhang, L., Sevinsky, C. J., Davis, B. M., Vertes, A., Anal Chem 2018, 90, 4626-4634.
19. Onjiko, R. M., Portero, E. P., Moody, S. A., Nemes, P., Anal Chem 2017.
20. Oomen, P., Aref, M. A., Kaya, I., Phan, N. T. N., Ewing, A. G., Anal Chem 2018.
21. Kempa, E. E., Hollywood, K. A., Smith, C. A., Barran, P. E., Analyst 2019.
22. Zhang, L., Vertes, A., Angew Chem Int Ed Engl 2018, 57, 4466-4477.
23. Zhang, W., Guled, F., Hankemeier, T., Ramautar, R., J Chromatogr B Analyt Technol Biomed Life Sci 2019, 1105, 10-14.
24. Moini, M., Anal Chem 2007, 79, 4241-4246.
25. Quist, J., Vulto, P., van der Linden, H., Hankemeier, T., Anal Chem 2012, 84, 9065-9071.
26. Raterink, R. J., Lindenburg, P. W., Vreeken, R. J., Hankemeier, T., Anal Chem 2013, 85, 7762-7768.
27. Drouin, N., Mandscheff, J. F., Rudaz, S., Schappler, J., Anal Chem 2017, 89, 6346-6350.
